MEIS1, a Promising Candidate Gene, Is Not Associated with the Core Symptoms of Antipsychotic-Induced Restless Legs Syndrome in Korean Schizophrenia Patients
Article information
Abstract
Objective
Restless legs syndrome (RLS) is a distressing sleep disorder to which individuals appear to be genetically predisposed. In the present study, we assumed that antipsychotic-induced RLS symptoms were attributable to differences in individual genetic susceptibility, and investigated whether MEIS1, a promising candidate gene, was associated with antipsychotic-induced RLS symptoms in schizophrenia patients.
Methods
All subjects were diagnosed with schizophrenia by board-certified psychiatrists using the Korean version of the Structured Clinical Interview for DSM-IV. We assessed antipsychotic-induced RLS symptoms in 190 Korean schizophrenic patients using the diagnostic criteria of the International Restless Legs Syndrome Study Group. Genotyping was performed for the rs2300478 and rs6710341 polymorphisms of the MEIS1 gene.
Results
We divided subjects into RLS symptom (n=96) and non-symptom (n=94) groups. There was no significant between-group difference in the genotype or allele frequencies of the two polymorphisms investigated, nor in the frequency of the rs2300478-rs6710341 haplotype.
Conclusion
Our data do not suggest that the rs2300478 and rs6710341 polymorphisms of the MEIS1 gene are associated with the core symptoms of antipsychotic-induced RLS in schizophrenia; different genetic mechanisms may underlie antipsychotic-induced vs. primary RLS.
INTRODUCTION
Restless legs syndrome (RLS), a sleep disorder characterized by uncomfortable sensations in, and irresistible urges to move, the legs, affects the quality of sleep and life. RLS is classified into primary and secondary types. Although the pathophysiology of primary RLS is not fully understood, some evidence suggests that it is caused by impaired central dopaminergic transmission and genetic factors. A family history of RLS has been reported in 60% of idiopathic RLS patients,1,2,3 and several genetic RLS loci (2q, 9p, 12q, 14q, 16p, 19p, 20p) have been identified using linkage analysis.4,5,6,7,8,9,10
Secondary RLS is caused by medical conditions including iron deficiency, chronic renal failure, neuropathy, pregnancy, and the use of medications such as antipsychotics and antidepressants.11 Several cases of antipsychotic-induced RLS have been reported,12,13,14 and the RLS incidence in schizophrenia patients is two-fold higher than in healthy controls, probably attributable to antipsychotic use.15
Although antipsychotics-induced RLS is generally regarded as being caused by dopamine receptor blockade, we suggest that this RLS may in fact be attributable to individual genetic susceptibilities, as is primary RLS. RLS does not occur in all patients using antipsychotics, some of whom show no symptoms at all. Therefore, we hypothesize that antipsychotic-induced RLS may be associated with differences in biological vulnerabilities to RLS, including genetic differences. To explore this hypothesis, we previously investigated associations between antipsychotic-induced RLS and several candidate genes [G protein β3 subunit (GNB3), tyrosine hydroxylase (TH), monoamine oxidase, dopamine receptor, BTB (POZ) domain-containing 9 (BTBD9) and circadian genes (i.e., the CLOCK and NPAS2 genes)]16,17,18,19 and demonstrated probable associations between RLS and the BTBD9, CLOCK, NPAS2, GNB3, and TH genes.16,19,20,21
In 2007, two independent genome-wide association studies (GWAS) were published, indicating a significant association between RLS and several genes not previously considered.22,23 Intronic variants of the myeloid ectropic viral insertion site 1 (MEIS1) and the homeobox 1 (MEIS1), mitogen-activated protein kinase 5 (MAP2K5), LBXCOR1 glyoxalase I (GLO1), BTBD9, and dynein axonemal heavy chain (DNAH8) gene, were strongly associated with RLS in the discovery phase of two GWAS.22,23 In the GWAS of Winkelmann et al., the rs2300478 polymorphism of the MEIS1 gene exhibited the strongest association with RLS (following Bonferroni correction) in a case/control sample.23 The haplotype consisting of rs6710341 allele A and rs12469063 allele G conferred the greatest risk of RLS.23 Several related and replication studies followed, investigating the association between the MEIS1 gene and the primary or secondary RLS caused by end-stage renal disease (ESRD).24,25,26 Therefore, the MEIS1 gene, which exhibited a positive association with both GWAS and secondary RLS caused by ESRD, is a good candidate for control of antipsychotics-induced RLS.
MEIS1, identified by Moskow et al.,27 is 137360 base-pairs in length and consists of 13 exons on chromosome 2p14.28 The MEIS1 protein has 390 amino acids and is expressed in murine hematopoietic stem cells, bone marrow, and fetal liver.29 Hematopoiesis and hepatic development were blocked in MEIS1-knockout mice.29,30 MEIS1 is also essential for limb formation in the mouse and chicken.31 The involvement of MEIS1 in limb formation and hematopoiesis may have implications for RLS development.
We investigated whether the MEIS1 gene was associated with the core symptoms of antipsychotic-induced RLS in schizophrenia.
METHODS
Subjects
A total of 190 schizophrenic patients were enrolled from Korea University Hospital and three other collaborating hospitals. All subjects were ethnic Koreans taking antipsychotics. They were diagnosed with schizophrenia by board-certified psychiatrists using the Korean version of the Structured Clinical Interview for DSM-IV.32 The exclusion criteria were as follows: 1) another Axis I diagnosis, a history of alcohol or other substance abuse, mental retardation, or a head injury or neurological disorder; 2) a serious medical illness or medical condition which could be mistaken for RLS; and 3) extreme psychotic symptoms preventing interview. The study was approved by the Ethics Committee of Korea University Medical Center, and all participants provided written informed consent.
Clinical assessments
RLS was assessed using the following four diagnostic criteria of the International Restless Legs Syndrome Study Group (IRLSSG):33 1) urge to move the legs, 2) unpleasant sensations in the legs, 3) urge to move or unpleasant sensations in the legs worsening during rest and relieved by bodily movements; and 4) urge to move the legs or unpleasant sensations that worsened or occurred only during the evening/nighttime. To obtain an adequate sample size (equalized across groups) for the genetic association study, we divided patients into the following two groups: a) subjects that met either criterion 1) or 2) of the IRLSSG diagnostic criteria (i.e., core symptoms of RLS); and b) subjects without RLS symptoms. RLS severity was determined using the IRLSSG rating scale for RLS.34 The Athens Insomnia Scale was used to assess insomnia severity,35 and the psychiatric symptoms of patients were evaluated using the Brief Psychiatric Rating Scale.36 The clinical characteristics of schizophrenia patients with antipsychotic-induced RLS have been previously described,15 and other findings on such subjects have been reported in previous studies.16,17,18,19,20,21
DNA analysis and genotyping
We selected the rs2300478 and rs6710341 polymorphisms of the MEIS1 gene for study, based on previous work, and the known minor allele frequencies in the general Korean population. Genomic DNA was extracted from whole blood using standard methods. Genotyping was performed via high-resolution melting (HRM) curve analysis. Polymerase chain reactions (PCRs) were performed, in a volume of 20 µL per reaction, in a 96-well Bio-Rad CFX96 Real Time PCR System (Bio-Rad, Hercules, CA, USA). The missing genotype rates for the rs2300478 and rs6710341 polymorphisms were both 0%.
Statistical analysis
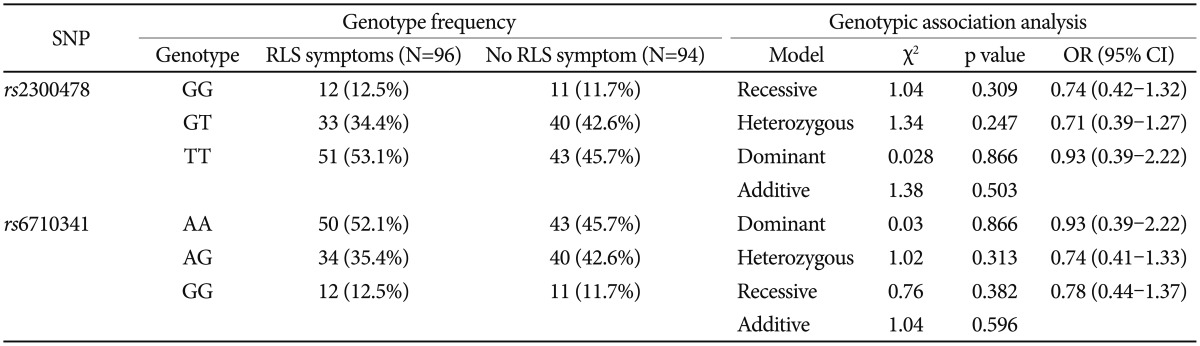
The presence of Hardy-Weinberg equilibrium was assessed using the χ2 goodness-of-fit test. Categorical data were analyzed using the χ2 or Fisher's exact test, and continuous variables were analyzed using Student's t-test or one-way analysis of variance. Haplotype frequencies were estimated using the SNPAlyze software package (ver. 7, DYNACOM, Chiba, Japan) based on use of an expectation-maximization algorithm and the maximum-likelihood approach. All analyses were performed using SPSS for Windows software (SPSS Inc., Chicago, IL, USA) and SNPAlyze. A value of p<0.05 was taken to indicate statistical significance, but after Bonferroni correction, the threshold was set at p<0.025 (Table 1 and 2), because two single-nucleotide polymorphisms were genotyped in the present study.
RESULTS
Patient mean age was 39.6±9.2 years (range: 22-66 years). Of 190 subjects, 84 (44.2%) were female and 106 (55.8%) male; 44 (23.2%) subjects met the IRLSSG diagnostic criteria and 96 (50.5%) had RLS symptoms. We divided subjects into with core RLS symptom (n=96) and non-symptom (n=94) groups; 52 of the core RLS symptom group did not meet all IRLSSG diagnostic criteria.
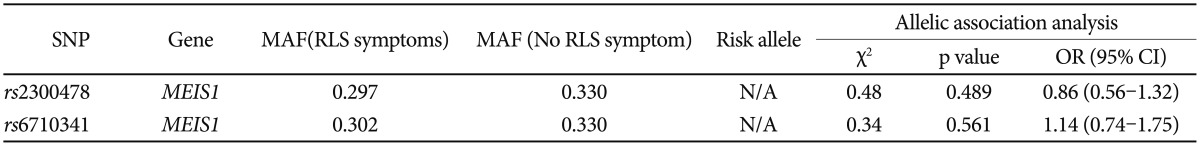
Genotype frequencies did not deviate from Hardy-Weinberg equilibrium (rs2300478; χ2=2.17, p=0.141; rs6710341; χ2=1.85, p=0.174). There was no significant group difference in genotype (rs2300478; χ2=1.38, p=0.503; rs6710341; χ2=1.04, p=0.596) or allele frequency (rs2300478; χ2=0.48, p=0.489; rs6710341; χ2=0.34, p=0.561) for either polymorphism investigated (Table 1 and 2).
Two markers (D'=0.9746, r2=0.9266, p=1.329×10-4) exhibited significant linkage disequilibrium (LD). There was no between-group difference in the frequency of the rs2300478-rs6710341 haplotype (Table 3).
DISCUSSION
Recently, several authors have reported cases of antipsychotic-induced RLS, a condition attracting increasing clinical attention. Symptoms of antipsychotic-induced RLS are commonly observed in schizophrenic patients taking antipsychotics; it is also notable that such patients exhibit more severe psychiatric symptoms than do those without RLS symptoms.15
However, the pathogenesis of antipsychotic-induced RLS remains poorly understood. Therefore, we suggest that genetic research on this topic is of value; idiopathic RLS has been considered to be a genetic disease in the previous GWAS and replication studies. In particular, MEIS1 is a promising candidate gene. Of the three gene regions, the rs2300478 polymorphism of the MEIS1 gene exhibited the strongest association with RLS during each stage of replication, and the three stages combined, in Winkelmann's GWAS.23 In addition, Xiong et al.24 demonstrated a significant association between rs2300478 and rs12469063, and RLS, in 28 patients and 140 control subjects, using autopsy brain samples.24 In two previous studies, MEIS1 played important roles in the development of proximo-distal limbs in animal studies,31,37 which could represent the underlying mechanism of the association between the MEIS1 gene and RLS.
However, in another GWAS study, using an Icelandic sample, there was no significant association between RLS and the MEIS1 gene region.22 A replication study in a European population investigated RLS loci including MEIS1 gene polymorphisms (rs6710341, rs12469063, rs2300478); there was no association between the MEIS1 gene and sporadic RLS, although a significant association was evident in a familial RLS case.25 In a genetic study of secondary RLS caused by ESRD, there was no significant association between the MEIS1 gene and RLS in the combined analysis, although there was an association in a German sample that was analyzed separately.26 A Taiwanese study of RLS caused by ESRD demonstrated a negative association with the MEIS1 gene.38 Such inconsistent data might be attributable to differences in genotype or allele frequencies among races. In fact, the rs6710341 genotyped in our study showed a minor allele frequency which is somewhat different from that in the past study.25 To validate this result, a replication study on the association of the MEIS1 gene with idiopathic RLS in a Korean sample is required. However, one study in which a relationship between MEIS1 variants and iron homeostasis was observed suggested that the pathophysiologies of idiopathic and antipsychotic-induced RLS differed.39
Our study had several limitations. First, the sample size may have been insufficiently large for results to be generalized. Second, the types of antipsychotics taken varied. Third, we used the presence or absence of core RLS symptoms to assign participants to groups, rather than the definitive RLS diagnostic criteria. Based on these limitations, the reported lack of association between the MEIS1 gene and RLS requires validation using a larger population, all of whom are taking identical antipsychotics. Future research should also evaluate the association between RLS and other candidate genes (for primary RLS; MAP2K5, LBXCOR1, GLO1, and DNAH8) reported elsewhere.22,23 Such studies would contribute considerably to our understanding of the pathogenesis of antipsychotic-induced and idiopathic RLS, and facilitate clinician selection of the safest antipsychotic for each patient.
Acknowledgments
This work was supported by the Gachon University research fund of 2012 (GCU-2012-M075).